Viral-Binding Protein Design Makes the Case for Intelligent Design Sick! (as in cool)
“What I cannot create, I do not understand.”
Richard Feynman
Nobody likes getting the flu. In fact, the influenza virus represents a serious health challenge. For most people it causes a few days of misery, but tragically for others, it takes their lives.
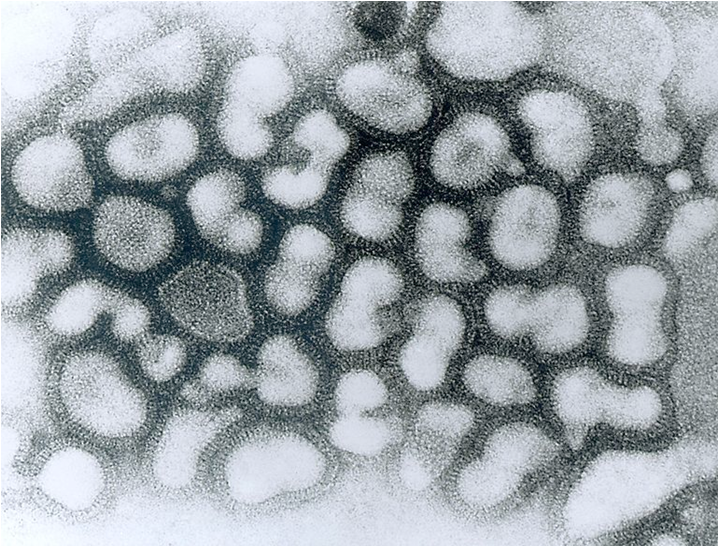
Image credit: Content Providers(s): CDC/ Dr. Erskine Palmer
Biomedical researchers are looking for ways to develop therapies against the strains of the influenza virus that avoid destruction by the immune system or fail to respond to antiviral drugs. Recently scientists from the University of Washington and The Scripps Institute made progress toward this end. They designed novel proteins from scratch that bind to the stem region of hemagglutinin, a viral protein that plays a central role in the infection process.1 The hope is that the new proteins will have therapeutic use and also help with diagnosis.
This advance has obvious biomedical importance. It also contributes toward understanding the fundamental nature of protein-protein interactions (PPIs), and with this insight comes powerful new evidence that life stems from a Creator.
Protein-Protein Interactions
Often, in order to carry out their function, proteins must interact and bind in a highly specific manner with other proteins. These PPIs are selective. If the wrong proteins bind to each other, the interaction is of no use to the cell.
A large and varied population of proteins crowds the cell’s interior. Even the simplest bacterium harbors several thousand different types of proteins, along with numerous copies of the other biomolecules inside it. The jam-packed environment complicates things. Proteins are more likely to encounter the “wrong” partners than the ones they are designed to interact with.
Biochemists are currently working to understand the specificity of PPIs and how proteins avoid unintended interactions. As it turns out, protein surfaces are carefully structured to allow strong interactions between protein pairs while minimizing the strength of the unwanted interactions. Recent work by Harvard scientists indicates that the concentration of PPI-participating proteins in the cell is also designed carefully.
In other words, protein structure and concentrations have to be precisely regulated to promote the PPIs critical for life. As I point out in my book The Cell’s Design, high-precision structures and interactions, exemplified by PPIs, are hallmark features of biochemical systems and, by analogy to fine-tuned human designs, point to the work of a Creator. The design of hemagglutinin-binding proteins by scientists further supports the idea that PPIs, and consequently biochemical systems, are the work of an intelligent Agent.
Hemagglutinin-Binding Proteins by Design
The University of Washington and Scripps Institute researchers gained insight into PPIs’ governing factors by designing from scratch proteins that interact with a designated protein in a specified manner. Part of the reason they chose to target hemagglutinin was to ensure that what they learned would not only contribute fundamental biochemical knowledge but also have immediate pragmatic use and biomedical application.
Methods exist to create proteins that can bind to other proteins. These approaches involve generating libraries of random proteins and then screening the library to identify the proteins that happen to bind to the target. The problems with this approach are: (1) it is impossible to control the location on the surface of the target protein to which the selected protein binds; and (2) it yields little fundamental understanding about the factors that dictate PPIs.
One way around these limitations is to design proteins from scratch with the desired binding properties. The University of Washington and The Scripps Institute researchers did this by first identifying the region of hemagglutinin they wanted the designed protein to bind to. They then determined what amino acids could bind strongly to the targeted region (called interaction hot spots) and how to arrange those amino acids in space to generate a shape complementary with the target site.
Once these steps were completed, the researchers needed to construct a protein that would possess the hemagglutinin-binding site after its backbone folded to adopt its three-dimensional shape. The researchers built the protein by searching through a set of 865 protein structures for scaffolds that would support the critical amino acids and position them in the appropriately spatial locations.
These procedures yielded 88 protein designs that would conceivably work. The researchers then built genes that encoded the information needed to make these proteins and cloned the genes into yeast. The yeast was genetically engineered to produce and display the designed proteins on their cell surface. This allowed the researchers to screen for protein designs that bind to hemagglutinin effectively.
Based on this process, the researchers identified two promising protein designs. They fine-tuned these structures to optimize binding by conducting in vitro evolution experiments. (See the video below for a discussion of in vitro evolution of proteins.)
Folge 37 – Directed Evolution of enantioselective enzymes from Chymiatrie on Vimeo.
Synthetic Biology and the Case for Intelligent Design
When considering this study, it is remarkable to note how much effort it took to design a protein that binds to a specific location on the hemagglutinin molecule. As biochemists Bryan Der and Brian Kuhlman point out while commenting on this work, the design of these proteins required:
…cutting-edge software developed by ~20 groups worldwide and 100,000 hours of highly parallel computing time. It also involved using a technique known as yeast display to screen candidate proteins and select those with high binding affinities, as well as x-ray crystallography to validate designs.2
If it takes this much work and intellectual input to create a single protein from scratch, is it really reasonable to think that undirected evolutionary processes could accomplish this task routinely?
In other words, the researchers from the University of Washington and The Scripps Institute have unwittingly provided empirical evidence that the high-precision interactions required for PPIs requires intelligent agency to arise. Sick!






