Why These 20 Amino Acids?
Biochemists have long puzzled over the 20 amino acids used to build proteins. Why these specific amino acids?
Over the last few decades, researchers have made some progress in addressing this question. Recent work from the University of Hawaii adds new insight, indicating that the set of amino acids used to make proteins is the optimal set. This discovery provides new evidence that life’s chemistry stems from the work of a Creator.
Kids love to ask “why” questions. Why is the sky blue? Why is grass green? Biochemists like asking why questions, too. One that has long fueled our curiosity relates to the choice of amino acids used to build proteins.
Amino acids are the building blocks that make up proteins. Twenty chemically distinct amino acids comprise the proteins found in every organism on Earth. That is, the set of amino acids used in biology is universal. Yet, hundreds of amino acids exist in nature. Biochemists want to know why the specific set of 20 amino acids, and not the others, occurs in proteins.
This question leads to other related why questions for biochemists:
- – Why are proteins built from amino acids? Why not build them from the chemically simpler hydroxy acids?
- – Why are the amino acids in proteins α-amino acids? Why not β– or γ– or δ-amino acids?
- – Why do all the amino acids in proteins have an α-hydrogen?
- – Why are there no N-alkyl amino acids in proteins?
Many naturally occurring amino acids possess these structural features. Shouldn’t at least some of these alternative compounds have made their way into proteins?
Over the course of the last 30 years or so, biochemists have sought answers to these questions.1 At first glance, it seems conceivable that other amino acids could have been chosen to fulfill the role of some, if not all, of the canonical amino acids.
But researchers from the University of Hawaii recently offered an insightful perspective on this scientific riddle that suggests otherwise. 2 The team conducted a quantitative comparison of the range of chemical and physical properties possessed by the 20 protein-building amino acids versus random sets of amino acids that could have been selected from early Earth’s hypothetical prebiotic soup. They concluded that the set of 20 amino acids is optimal.
It turns out that the set of amino acids found in biological systems possess properties that evenly and uniformly varies across a broad range of sizes, charges, and hydrophobicities. They also demonstrate that the amino acids selected for proteins is a “highly unusual set of 20 amino acids; a maximum of 0.03% random sets out-performed the standard amino acid alphabet in two properties, while no single random set exhibited greater coverage in all three properties simultaneously.”3
This amino acid set’s wide range of physicochemical properties make it possible for proteins to carry out critical chemical operations necessary for life. In my view, this insight supports the notion that life’s chemistry has been designed through the work of an intelligent Agent.
Answers to the “Why” Questions
Even prior to this most recent study, biochemists had the sense that an exquisite chemical logic undergirds the choice of amino acids that make up proteinsα. What follows is a quick survey of some of the insights that have been accumulated over the past thirty years, or so.
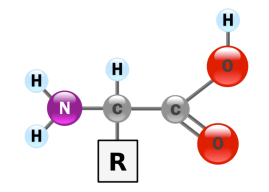
Figure 1. The structure of an α-amino acid. Image credit: YassineMrabet.
Why not hydroxy acids? Just as amino acids are linked to make proteins, hydroxy acids can be linked together in a head-to-tail fashion to form molecular chains. However, proteins cannot be built from hydroxy acids because the bond that joins these molecules together consists of an ester linkage, not an amide linkage as is found in proteins. Ester linkages are not as chemically stable as amide linkages. (They are susceptible to basic hydrolysis.) Additionally, ester linkages do not form a planar bond between subunits. On the other hand, amide bonds are planar, which allows intrastrand hydrogen bonding. These bonds stabilize the higher order protein structures that dictate the function of these biomolecules.
Amide bonds could be generated if the subunit molecules used to make up the protein chains alternated between diacids and diamines. This type of bonding occurs in nylons, molecular analogs to proteins. However, the use of these subunits introduces unnecessary chemical complexity into the system without any additional payoff. In other words, amino acids generate amide bonds in the simplest, most efficient way possible.
Why α-amino acids.? Unlike α-amino acids, β– or γ– or δ-amino acids can’t form orderly secondary structures—like α-helicies or β-pleated sheets—critical for protein structure. N-alkyl amino acids are also unsuitable as protein subunits because they cannot participate in hydrogen bond interactions.
Additionally, the α-amino acids must possess a hydrogen atom in the α position. This helps avoid steric hindrance when the protein folds into higher order three-dimensional structures.
The R Groups
The α-amino acids found in proteins have another important structural feature called an R group. This feature varies from amino acid to amino acid and invests these compounds with distinct chemical and physical properties.
Biochemists have some understanding about these R groups’ physicochemical utility. It turns out that, thanks to the R groups, the relatively small set of 20 amino acids possesses a wide range of chemical and physical properties, which imparts a significantly diverse array of structural and functional possibilities to proteins.
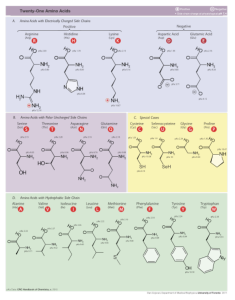
Figure 2. The side chains of the 20 protein-building amino acids. Image credit: Dancojocari.
Hydrophobic groups. For example, some of the R groups are hydrophobic, seeking to avoid contact with water at all costs. These amino acids are often buried in the protein interior where they play a key role in stabilizing the protein’s three-dimensional structure. It turns out that the R groups of several of the hydrophobic amino acids consist of branched alkyl chains. This leads to reduced mobility within the protein interior and holds together the protein structure. This benefit would not exist if the R groups consisted of linear, unbranched chains.
The R group of glycine is a hydrogen atom. This, too, has steric benefits for protein structure, allowing the protein chain to make tight turns where glycine residue occurs.
Hydrophilic groups. Other R groups are hydrophilic. Again, it appears as if these amino acids have been carefully selected for their diversity of useful physicochemical properties.
Alcohols. The amino acids serine and threonine are the simplest primary and secondary alcohols possible. Alcohols are useful for a number of chemical processes. They serve as attachment points for chemical groups like phosphates (which modify protein structure and function) and take part in a number of important chemical reactions.
Thiol groups. The amino acid cysteine is the simplest thiol R group possible. Thiols can take part in important chemical reactions as well. In addition, the R group of one cysteine residue in a protein can react with the R group of another cysteine to form a crosslink that helps stabilize protein structure.
Carboxylic acid groups. Aspartic acid and glutamic acid provide proteins with carboxylic acid groups, which give proteins the capacity for acid/base chemistry. When the carboxyl group is ionized, these amino acids become negatively charged and can interact with amino acid groups bearing a positive charge to form a salt bridge. These molecular-scale “bridges” help stabilize protein structure. Interestingly, the R group of glutamic acid is longer than that of aspartic acid. This difference allow for salt bridges of varying lengths to form within a protein’s interior.
Positively charged groups. Lysine, histidine, and arganine are the three positively charged amino acids. The R group of arganine bears a permanent positive charge. On the other hand, the positive charge of the lysine and histidine R groups depends on the specific chemical environment. In other words, the positive charge of these side groups can be “turned on and off.”
Lysine’s positive charge stems from the presence of an amino group while histidine’s positive charge is a property of the imidazole functional group. Not only do these two R groups provide a source of positive charge, but they also have distinct chemical properties that add to the range of protein functional properties.
Aromatic hydrocarbons. Phenylalanine, tyrosine, and tryptophan all have R groups made up of aromatic hydrocarbons. Phenylalanine possesses the simplest possible R group structure that can contain a benzene ring. The benzene ring is hydrophobic and planar, making it structurally rigid. Additionally, the electron delocalization within the aromatic ring imparts phenylalanine with the capacity to take part in chemical interactions where typical hydrophobic R groups cannot. Likewise, tyrosine’s phenolic R group gives this aromatic amino acid the wide range of interesting chemical properties possessed by phenol.
Technically, given the chemical nature of its R group, proline is not an amino acid. Rather it is an imino acid. Proline is an extremely rigid molecule. When proline is incorporated into a protein chain, its R group takes part in forming the backbone of the protein chain. This forces the protein chain to take a sharp turn that proline’s structural rigidity lock’s into place.
Of note are the functional R groups absent from the 20 protein-forming amino acids, particularly aldehydes and ketones. These two chemical functionalities take part in a wide range of interesting chemistry, which, on one hand, makes their omission from the set of 20 a little puzzling. On the other hand, these two functional groups are so reactive that their presence in a protein molecule would be disruptive. For example, both groups can form chemical crosslinks. This reaction could easily (and randomly) tether two protein molecules together. Unwanted crosslinks would form within a protein as well.
Biochemical Optimization and the Case for Intelligent Design
Evolutionary biologists argue that the undirected processes of chemical and natural selection generated the set amino acids used to make proteins. If this is the case, some level of optimization would be expected, but not the extreme optimization just discovered by the University of Hawaii researchers.
The widespread expectation is that evolutionary mechanisms should produce systems that work “just good enough” for the organism to survive, but are not necessarily as optimized as the set of 20 protein-forming amino acids. Yet the amino acids found in proteins do not look the way one would expect based on evolutionary models.
First of all, it is suspicious that the amino acids found in proteins don’t contain α-amino-n-butyric acid, norvaline, and norleucine. According to evolutionary models for the origin of life, these amino acids should have occurred at extremely high levels on early Earth. They would have been a readily available source of amino acids for the first proteins. Additionally, they possess physicochemical properties similar to those of the hydrophobic amino acids found in proteins. Second, the existence of asparagine and glutamine in proteins is intriguing. Scientists do not believe that these amino acids existed on the primordial Earth.
As an alternative explanation for the optimized set of amino acids found in proteins, I propose the work of a Creator. As I discussed in my book The Cell’s Design, objects and systems created and produced by human designers are typically optimized. Optimization is an indicator of intelligent design, achieved through foresight and preplanning. It requires inordinate attention to detail and careful craftsmanship. By analogy, the optimized biochemistry, epitomized by the amino acid set that makes up proteins, could be rationally understood as the work of a Creator.
Endnotes
- Arthur L. Weber and Stanley L. Miller, “Reasons for the Occurrence of the Twenty Coded Protein Amino Acids,” Journal of Molecular Evolution 17, no. 5 (1981): 273–84; H. James Cleaves II, “The Origin of the Biologically Coded Amino Acids,” Journal of Theoretical Biology 263, no. 4 (2010): 490–98.
- Gayle K. Philip and Stephen J. Freeland, “Did Evolution Select a Nonrandom ‘Alphabet’ of Amino Acids?” Astrobiology 11 (April 2011), 235–40.
- Ibid.